Modifications
General Approach
Most protein samples exhibit some degree of
modification.
There are the "natural" post translational modifications, such as phosphorylation
and glycosylation. There are the accidental modifications which
are artefacts of sample handling, such as oxidation. Finally,
there are the modifications deliberately introduced during sample
work-up, such as cysteine derivatisation. In most cases, it is
only the deliberate modifications which are known about for certain
at the time of doing a search.
It might be assumed that the search software could allow for
those modifications which are described in sequence entry annotations.
However, writing code to parse these sequence annotations would
be a major task. Indeed, many post-translational modifications
are not specified in a way which can be readily translated into
specific mass differences. For example, noting that a residue
is an actual or potential glycosylation site is not much help.
Even a simple modification, such as phosphorylation, is rarely
quantitative, so that it would be necessary to include mass values
for all permutations of occupied and unoccupied sites.
And, of course, protein sequences derived translated from nucleotide
sequences contain no information on post translational modifications.
The solution adopted here is to allow modifications to be specified
in two different ways: fixed modifications and variable modifications.
Fixed modifications are applied
universally, to every instance of the specified residue or terminus. There is
no computational overhead associated with a fixed modification, it is simply
equivalent to using a different mass for the modified residue or terminus.
For example, selecting Carboxymethyl (C) means that all calculations will
use 161 Da as the mass of cysteine.
Variable modifications are those which may or may not be present. Mascot
tests all possible arrangements of variable modifications to find the best match.
For example, if Oxidation (M) is selected, and a peptide contains 3
methionines, Mascot will test for a match with the experimental data for that
peptide containing 0, 1, 2, or 3 oxidised methionine residues. This greatly
increases the complexity of a search, resulting in longer search times and
reduced specificity, so variable modifications should be used sparingly.
(Quantitation methods support an additional mode:
Exclusive modifications.)
Unimod
The list of modifications used by Mascot is taken directly from the
Unimod database. For further
details of individual modifications, please refer to Unimod. Note that Unimod
is a community supported resource. If you want to add a new modification
to Unimod, you can do so, and you then become the curator of the new record. The Mascot
modifications list on the public web site is updated from Unimod each weekend.
By default, only selected modifications are displayed in the
Mascot search form. If you want to see the complete list, you must go to the
search form defaults page and tick the
checkbox for 'Show all mods.'.
In Mascot 2.1 and earlier, modification definitions were stored in a
configuration file called mod_file. Mascot now takes its modification definitions direct from an XML representation of the
Unimod database. To update the local
definitions, simply download the latest XML file from the Unimod
help page.
In Unimod, both amino acid residues and modifications are defined in terms of their elemental
composition. This is important for metabolic labelling, in which the isotopic
label is present throughout the peptide backbone. If you want to view or edit the local
unimod.xml file, a browser-based Configuration Editor is provided:
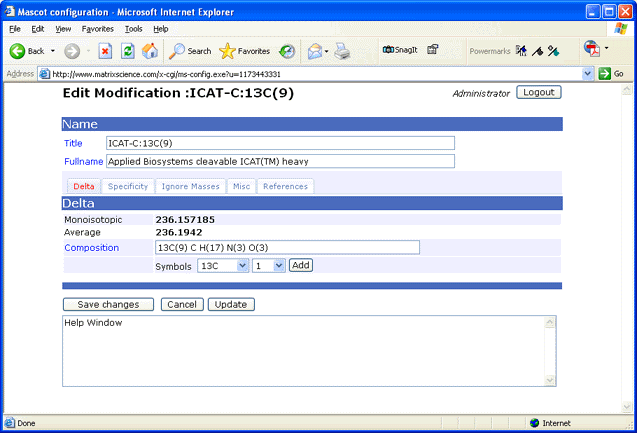
Note: Whenever unimod.xml is updated, an equivalent mod_file is created automatically to support
old client applications that require this file.
Do not be tempted to edit mod_file, because any changes will be lost the next time
unimod.xml is updated.
Other lists of modifications
DeltaMass is a comprehensive
list of modifications, sorted by mass.
RESID database contains detailed descriptions of many
post-translational modifications.
Phosphorylation is one of the most interesting and studied modifications.
It is also one
of the most challenging for database searching, because of these factors:
- Site heterogeneity
- 3 fragmentation channels
- intact fragments
- neutral loss of HPO3 (80 Da)
- neutral loss of H3PO4 (98 Da)
- Can occur at STY - ~16% of residues.
Support for a single neutral loss per modification was introduced in Mascot 1.7.
Mascot 2.1 added support for multiple neutral losses from both
fragment ions and the precursor.
In Mascot 2.2, the default phosphorylation modifications, that appear on the "short"
modifications list, appear in the mod_file as:
Title:Phospho (ST)
Residues:S 166.998359 167.0572
Residues:T 181.014010 181.0838
NeutralLoss:97.976896 97.9952
NeutralLoss:0 0 0
*
Title:Phospho (Y)
Residues:Y 243.029660 243.1532
*
That is, pY always stays intact,
while pS and pT can stay intact or can lose 98.
This is not a hard and fast rule, and sometimes a
loss of 80 is also observed. However, this is not included in the
definition because it is
identical to the delta of the original modification.
Allowing for the possibility of 80 Da neutral loss introduces ambiguity as to
the site of the modification when there are multiple potential phosphorylation
sites in a peptide. For example, this match to pTESPATAAETASEELDNR
gets a score of 115
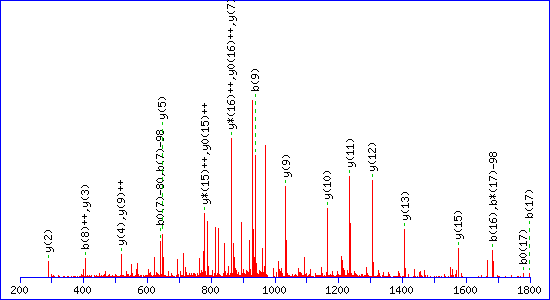
If a neutral loss of 80 Da is allowed, the score
for a match to TESPATAAETApSEELDNR is almost as high, 92
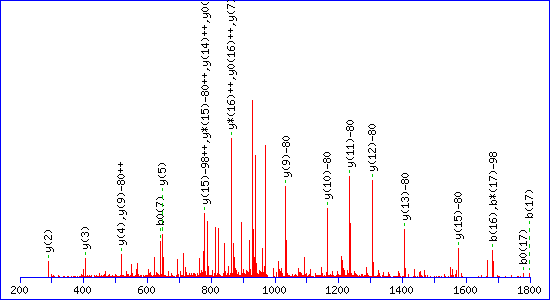
The reason is clear. The matching peaks are all y ions, so the point of
modification can be shifted towards the C-terminus by swapping the matching
series from y to y-80. Without the availability of an 80 Da loss, the score for the
second match drops to 29.
It has often been observed that the neutral loss from the precursor can be an
excellent guide to the identity of the phosphorylated residue. If a strong loss
of 98 Da is observed, then the expectation is pS or pT.
If no neutral loss, then pY. In Mascot, one or more precursor neutral losses can be specified. They
can also be made "required", which means that the peak
must be present in the spectrum. This carries some risk, because a
perfectly good match could be rejected if this peak happened to be missing.
|